Frontiers | DMfold: A Novel Method to Predict RNA Secondary Structure With Pseudoknots Based on Deep Learning and Improved Base Pair Maximization Principle
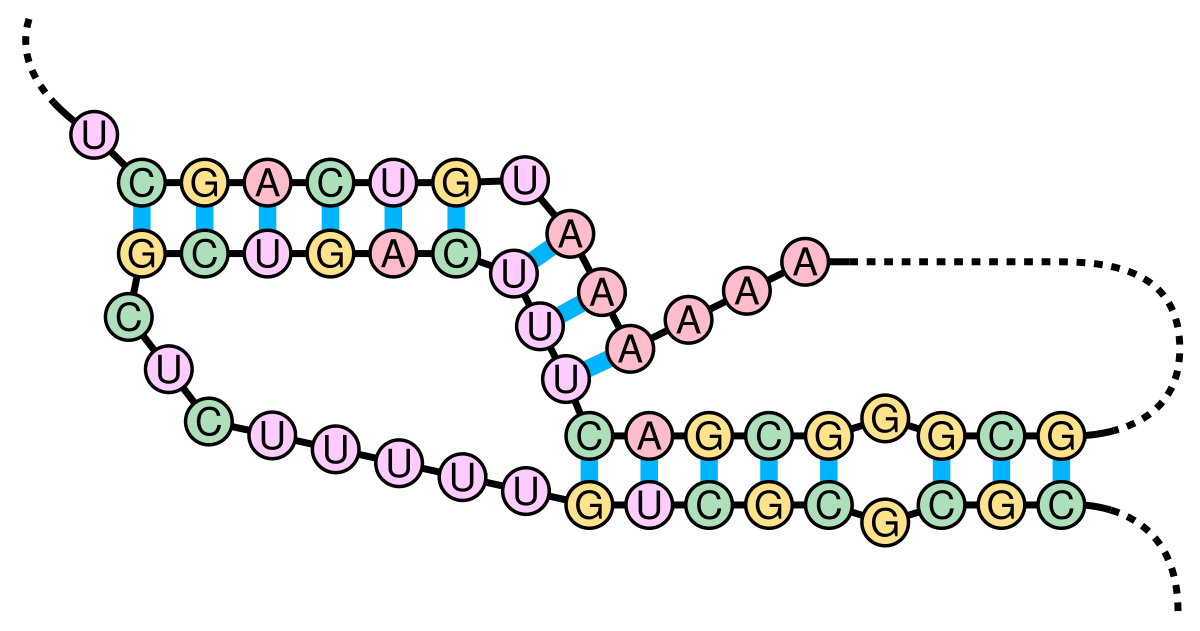
RNA structure types. (A) cartoon schematic of RNA structure types. (B)... | Download Scientific Diagram

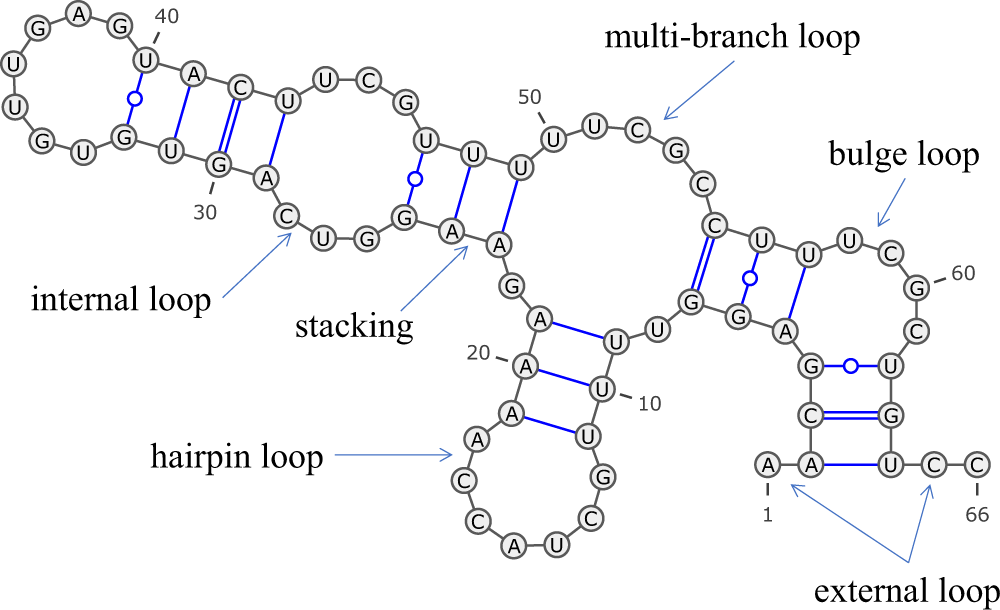
An RNA secondary structure illustrating the types of features included... | Download Scientific Diagram

Multistrand RNA Secondary Structure Prediction and Nanostructure Design Including Pseudoknots | ACS Nano

Structures of artificially designed discrete RNA nanoarchitectures at near-atomic resolution | Science Advances

PDF) Linear Time Algorithm for Calculating the Free Energy of the RNA Secondary Structure Including Pseudoknots (VS other algorithms) | Mohammad Ali Safari - Academia.edu

A partition function algorithm for nucleic acid secondary structure including pseudoknots - Dirks - 2003 - Journal of Computational Chemistry - Wiley Online Library

Small molecule targeting of biologically relevant RNA tertiary and quaternary structures - ScienceDirect
RNA secondary structure prediction using deep learning with thermodynamic integration | Nature Communications
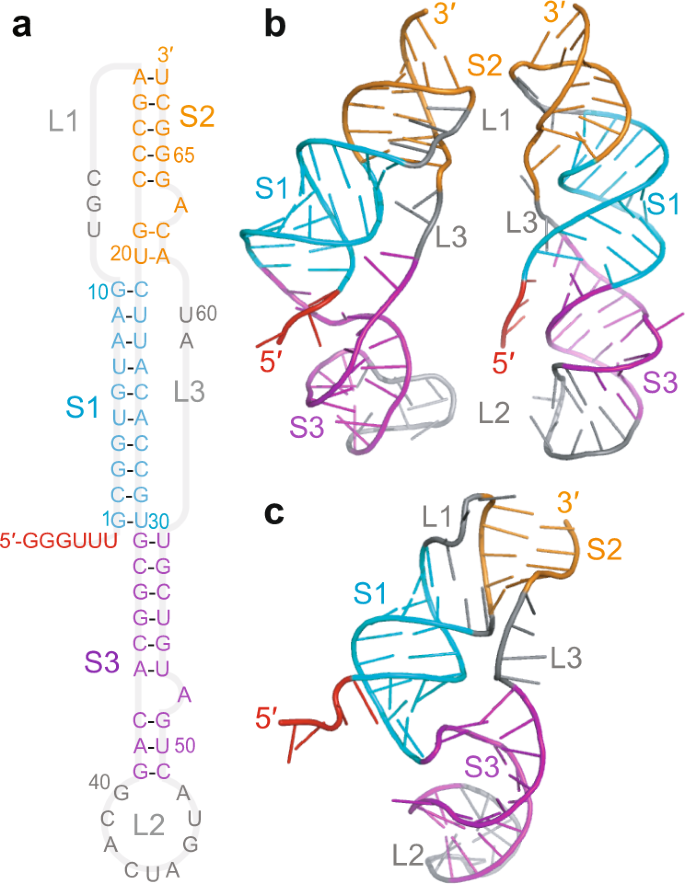
Structural dynamics of single SARS-CoV-2 pseudoknot molecules reveal topologically distinct conformers | Nature Communications

Structural dynamics of the SARS-CoV-2 frameshift-stimulatory pseudoknot reveal topologically distinct conformers | bioRxiv
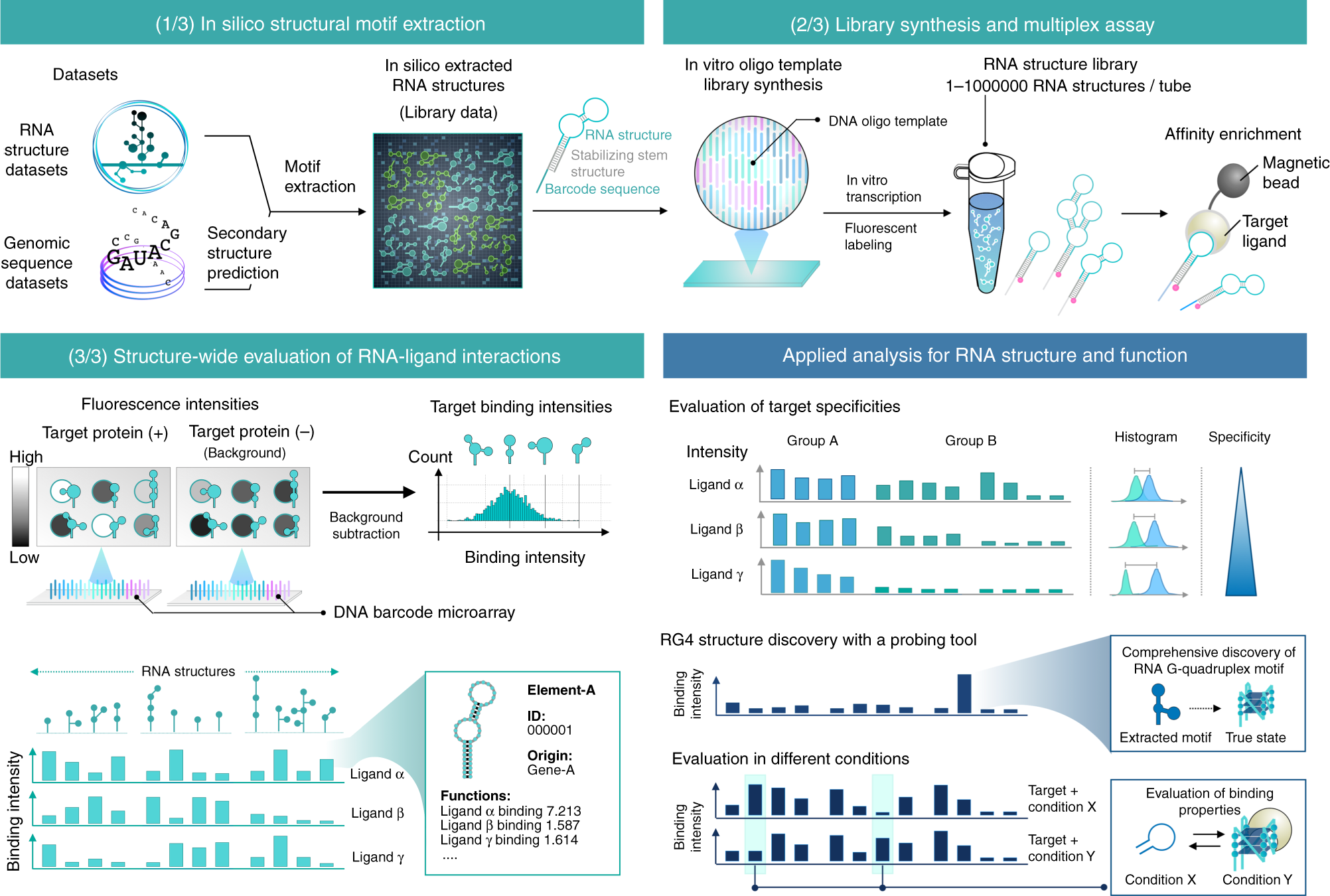
RNA structure-wide discovery of functional interactions with multiplexed RNA motif library | Nature Communications
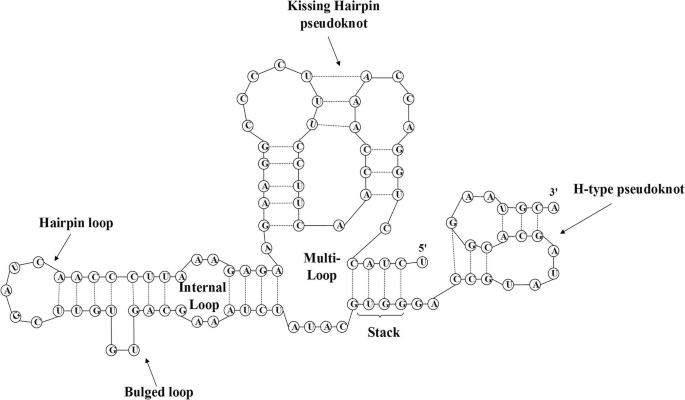
An efficient simulated annealing algorithm for the RNA secondary structure prediction with Pseudoknots | BMC Genomics | Full Text
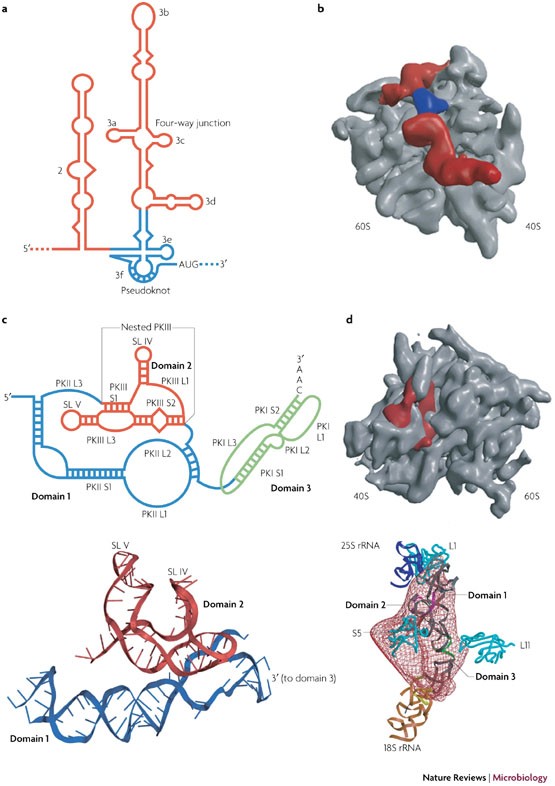
Viral RNA pseudoknots: versatile motifs in gene expression and replication | Nature Reviews Microbiology

Structural elements: the five structural elements of RNA structure,... | Download Scientific Diagram
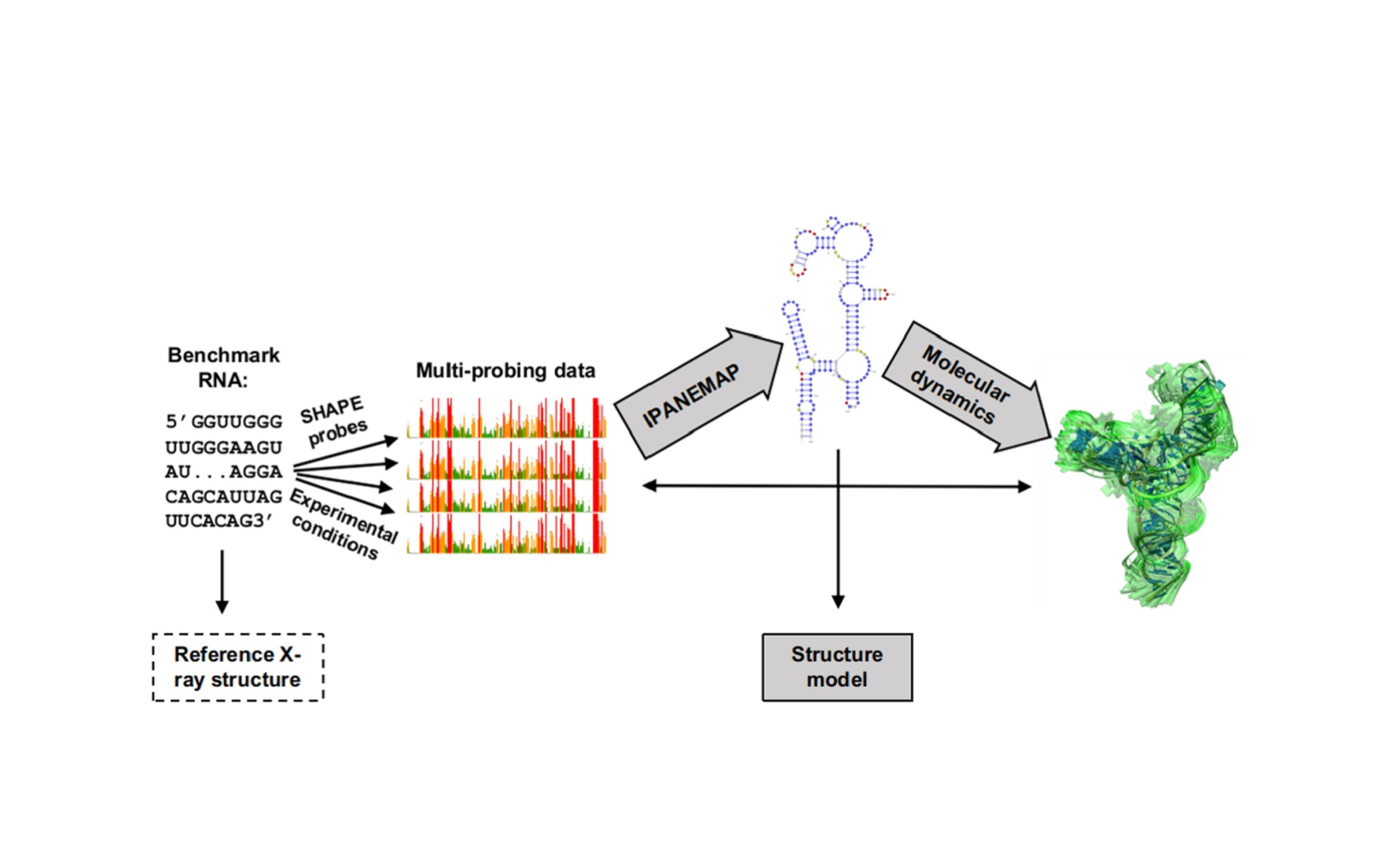
ncRNA | Free Full-Text | Progress toward SHAPE Constrained Computational Prediction of Tertiary Interactions in RNA Structure

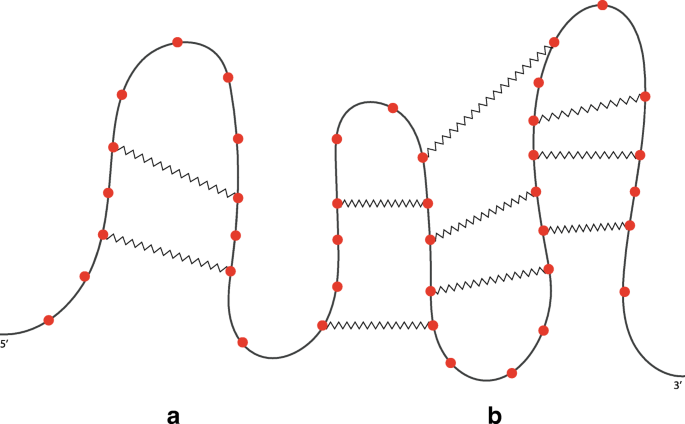
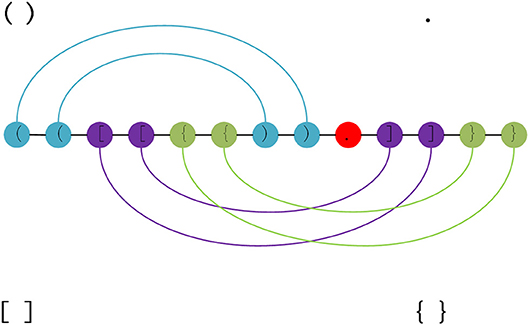
Pseudoknotted and pseudoknot-free secondary structures. Examples of... | Download Scientific Diagram