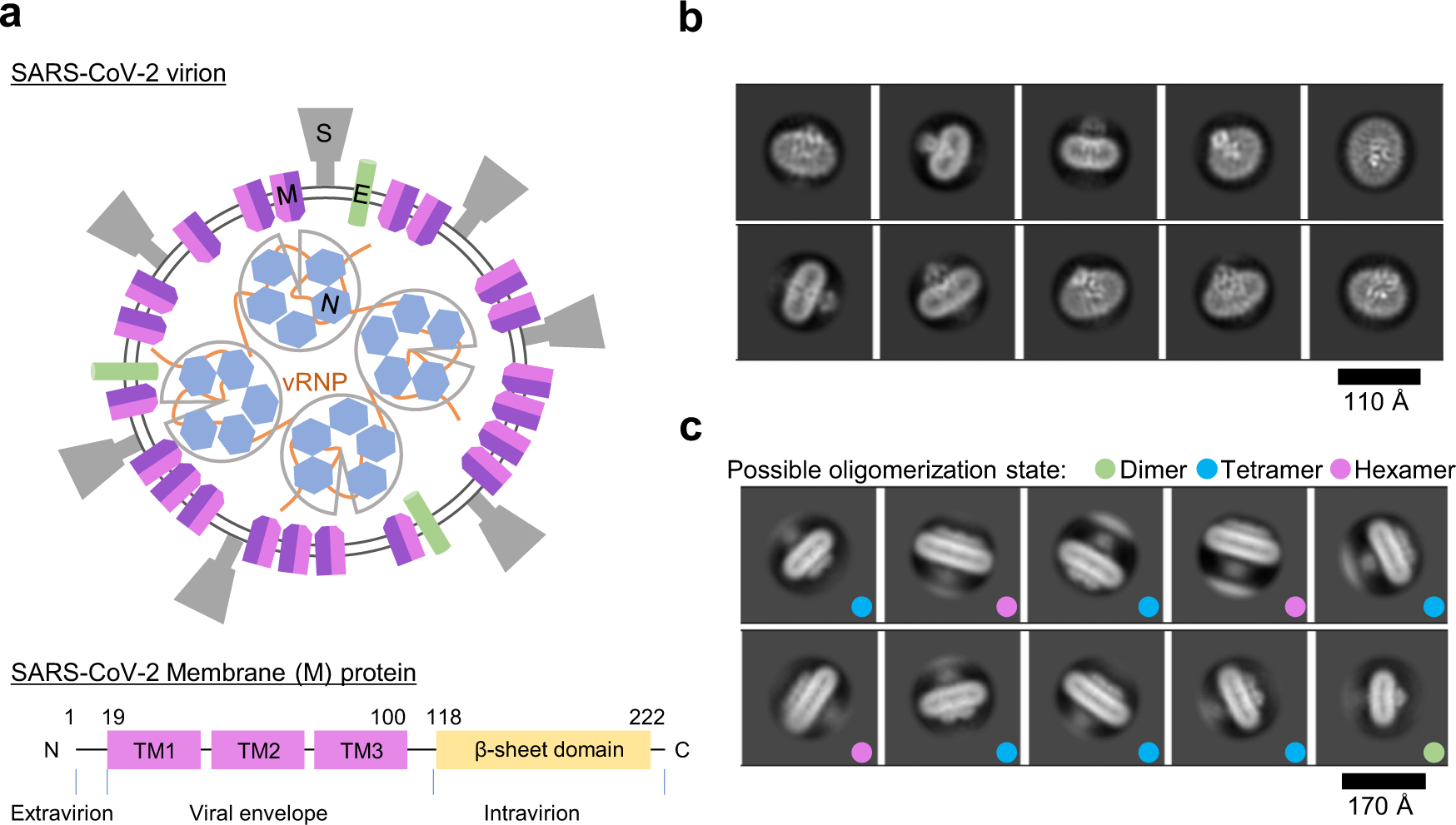
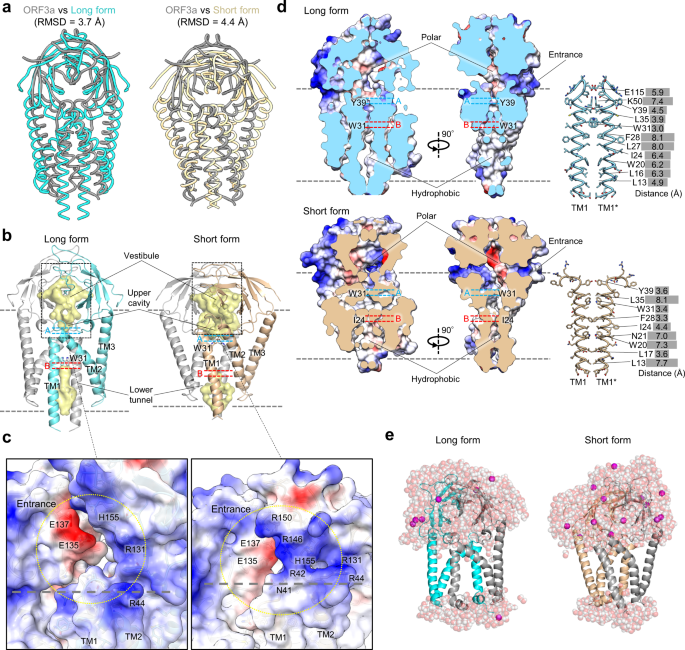
Probing the formation, structure and free energy relationships of M protein dimers of SARS-CoV-2 - ScienceDirect

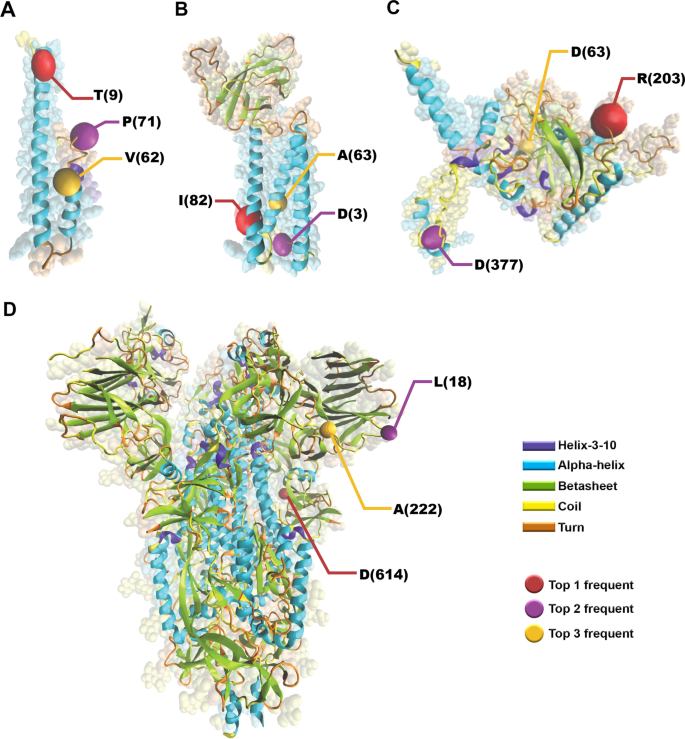
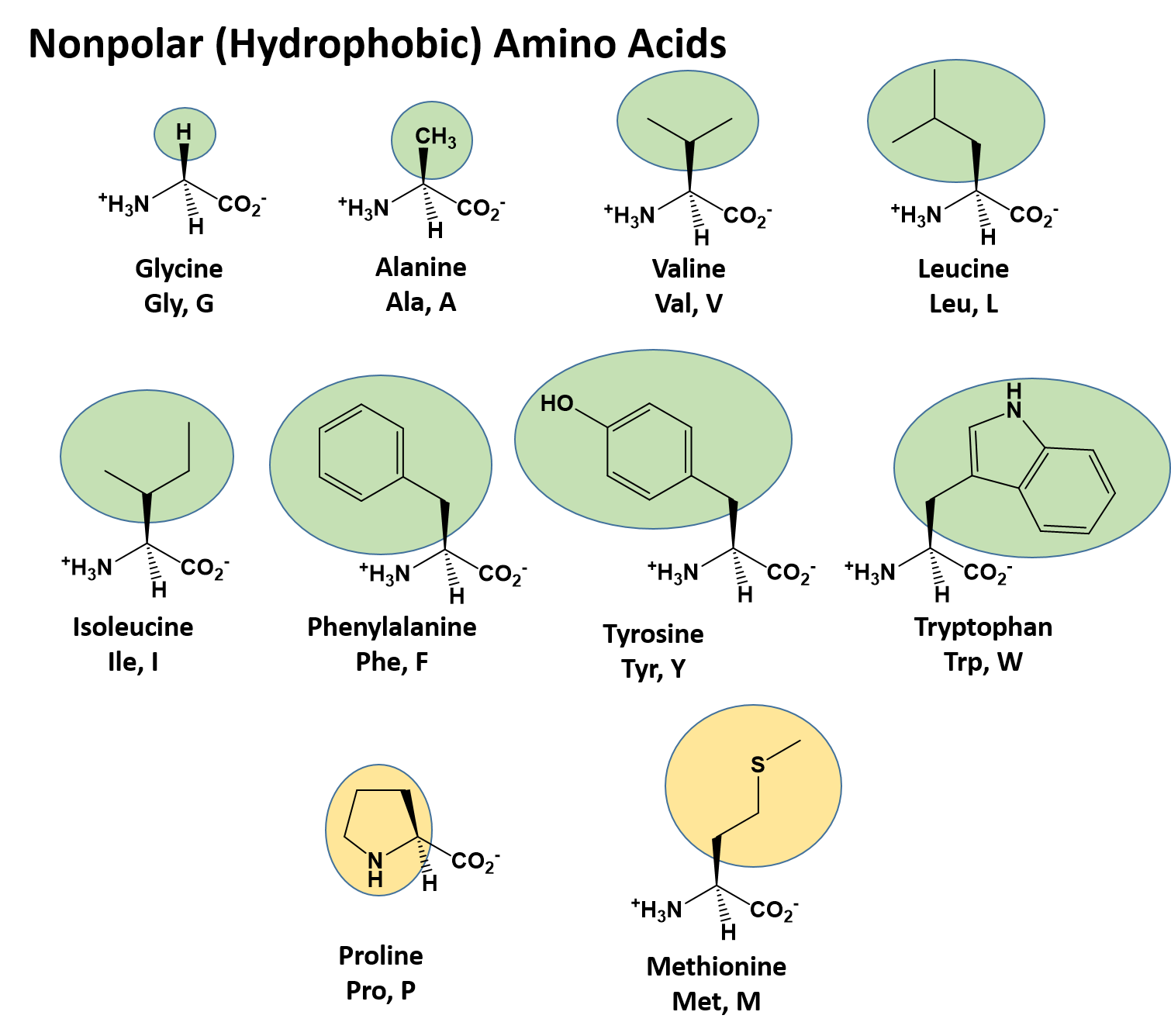
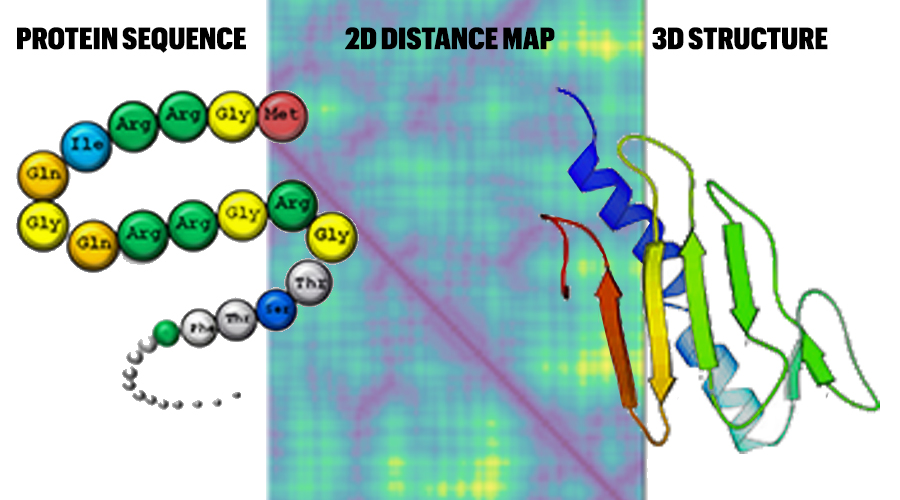
Protein structure, amino acid composition and sequence determine proteome vulnerability to oxidation‐induced damage | The EMBO Journal

Architecture and self‐assembly of the SARS‐CoV‐2 nucleocapsid protein - Ye - 2020 - Protein Science - Wiley Online Library

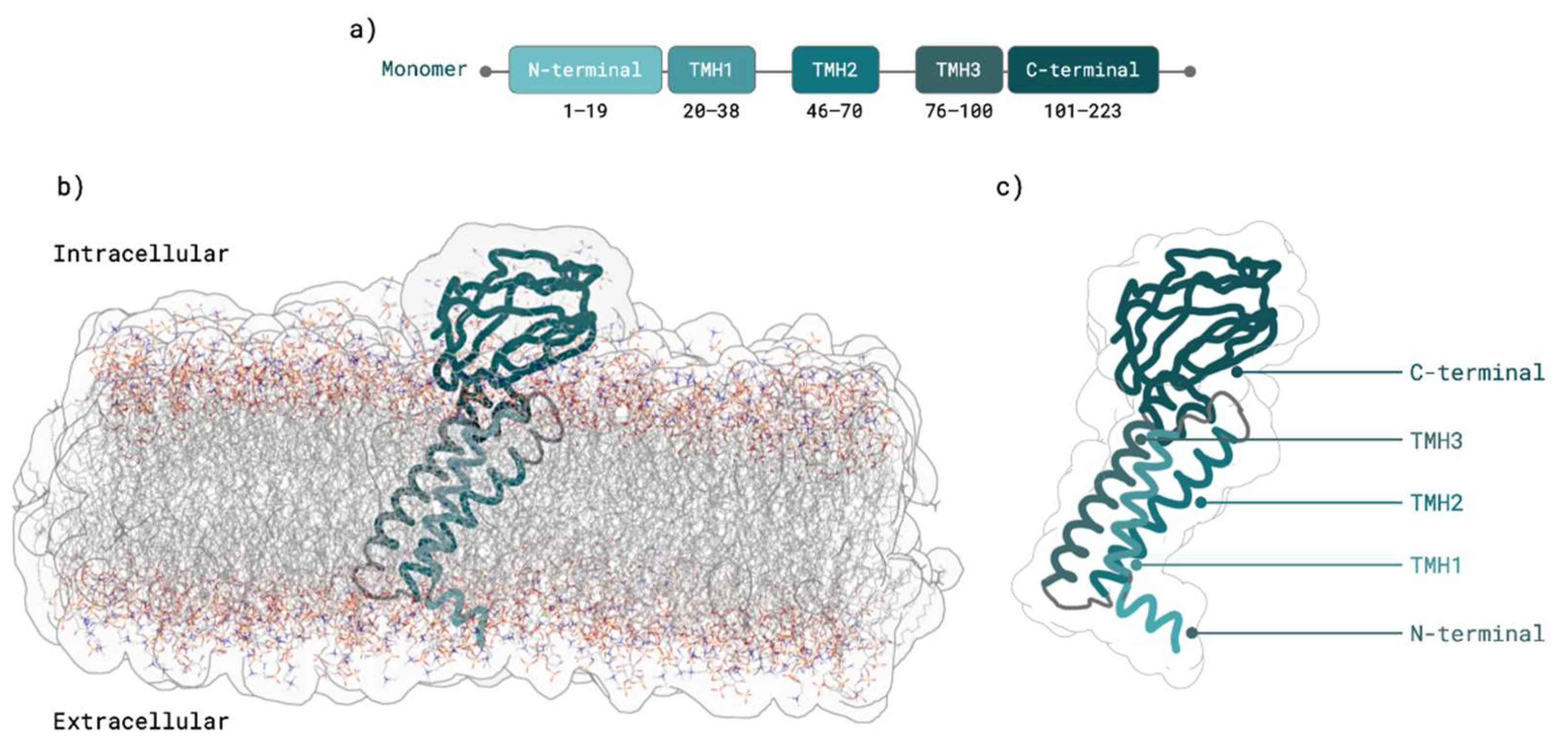
Structure and dynamics of the SARS‐CoV‐2 envelope protein monomer - Kuzmin - 2022 - Proteins: Structure, Function, and Bioinformatics - Wiley Online Library

Structural determination of Streptococcus pyogenes M1 protein interactions with human immunoglobulin G using integrative structural biology | bioRxiv
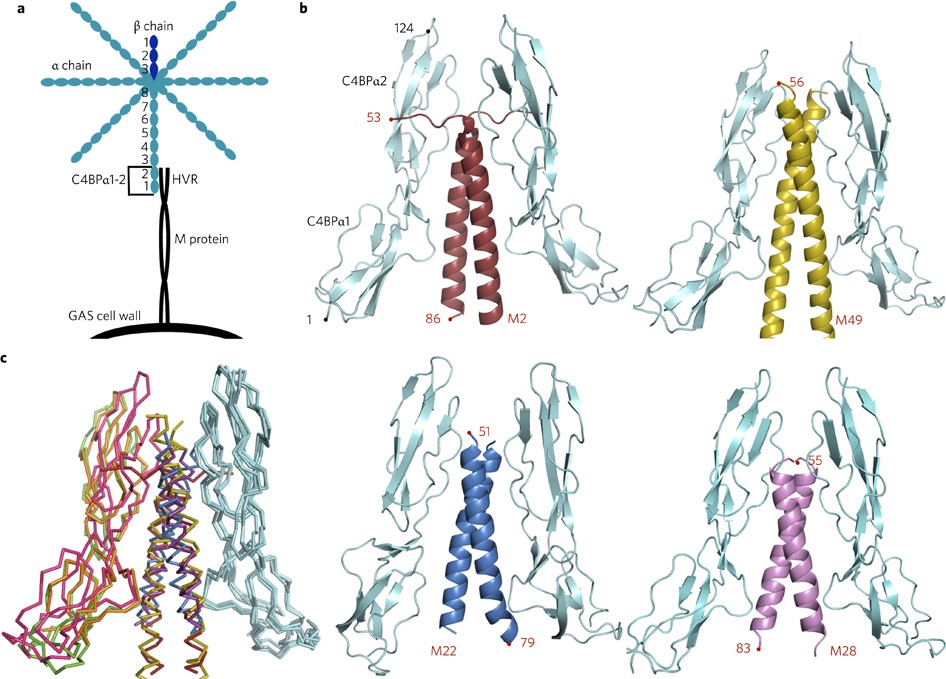
Conserved patterns hidden within group A Streptococcus M protein hypervariability recognize human C4b-binding protein | Nature Microbiology

Structure and Function of Major SARS-CoV-2 and SARS-CoV Proteins - Ritesh Gorkhali, Prashanna Koirala, Sadikshya Rijal, Ashmita Mainali, Adesh Baral, Hitesh Kumar Bhattarai, 2021